Research
The Champion Lab is a highly collaborative group interested in applying analytical instrumentation and techniques to answer biological questions. Our areas of expertise include bottom-up proteomics, mass spectrometry, and capillary separations. We apply these techniques to our model organisms in the genus Mycobacterium, including the causative agent of tuberculosis.
|
Bottom-Up Protemics
|
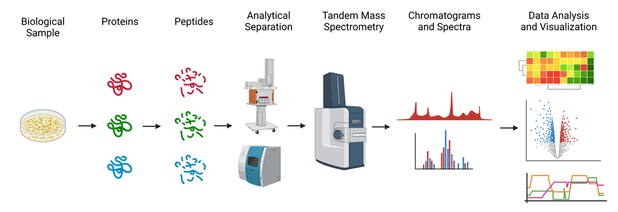
We use mass spectrometry for bottom-up proteomics (BUP), a versatile technique to identify and quantify peptides and proteins in samples. We develop novel strategies for all steps of the proteomics process: sample preparation and labeling, chromatographic and electrophoretic separations, mass spectrometry instrumentation, and informatics-based data analysis and data visualization. Check out the sections below to see details on our projects in all of these areas.
|
Sample Preparation
Robotics and Automation |
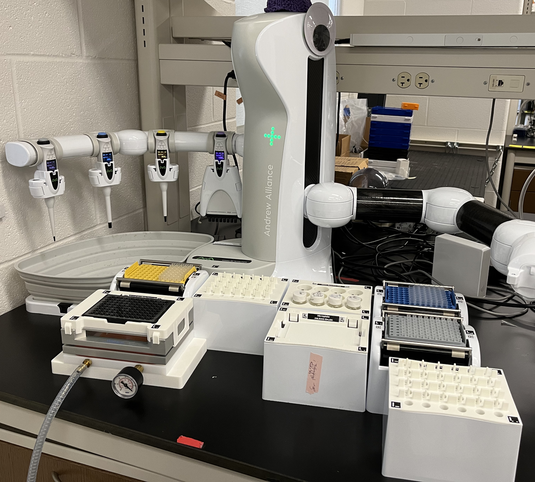
Identifying and quantifying changes in protein abundance is central to the understanding of physiological processes. The major technique used for interrogating complex protein mixtures is bottom-up proteomics. Throughput of this process is surprisingly low; a core facility proteomics lab might analyze 20 samples/day per instrument using nano-liquid chromatography with tandem mass spectrometry (nLC-MS/MS). As a consequence of this, automation of proteomics is rare, and virtually all preparation is performed by hand. This leads to decreased accuracy and precision between samples and labs.
We have adapted a lower-cost robot arm to perform proteomics preparation for meso-scale throughput. We altered a commercial lab robot, an Andrew+ to automate these processes (Shown to the left). Sample preparation in bottom-up proteomics has two broad steps; digestion and desalting. We adapted the most popular and common techniques used in proteomics for automation. In-solution/S-Trap digestion, ZipTip desalting, SPE desalting and MALDI based preparation. Custom bridges and adapters produced on consumer 3D printers facilitated rapid development of custom applications and utility. |
|
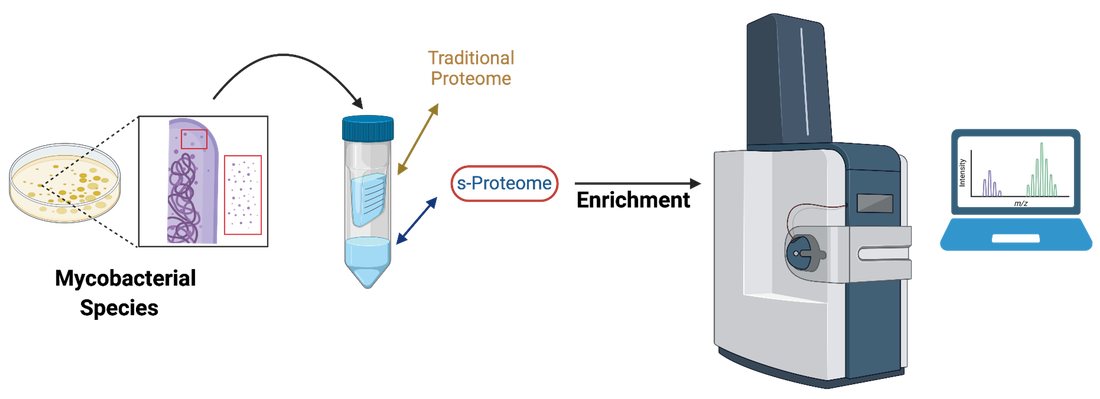
Small proteins (sProts) are typically defined as containing fewer than 100 amino acids and are inherently difficult to identify from genomics and directly by mass spectrometry. sProts are a widely abundant class of protein encoded in all forms of life yet remain unannotated and understudied. Previous research shows that the few sProts that have been functionally characterized demonstrate distinct cellular roles. The lack of annotation of sProts is because traditional sample preparation methods are biased toward high molecular weight proteins and discard most of the small proteome. We aim to fill this knowledge gap by developing new methods to enrich and functionally characterize the small proteomes of mycobacteria.
|
Small Proteins
|
N-terminomics and Labeling |
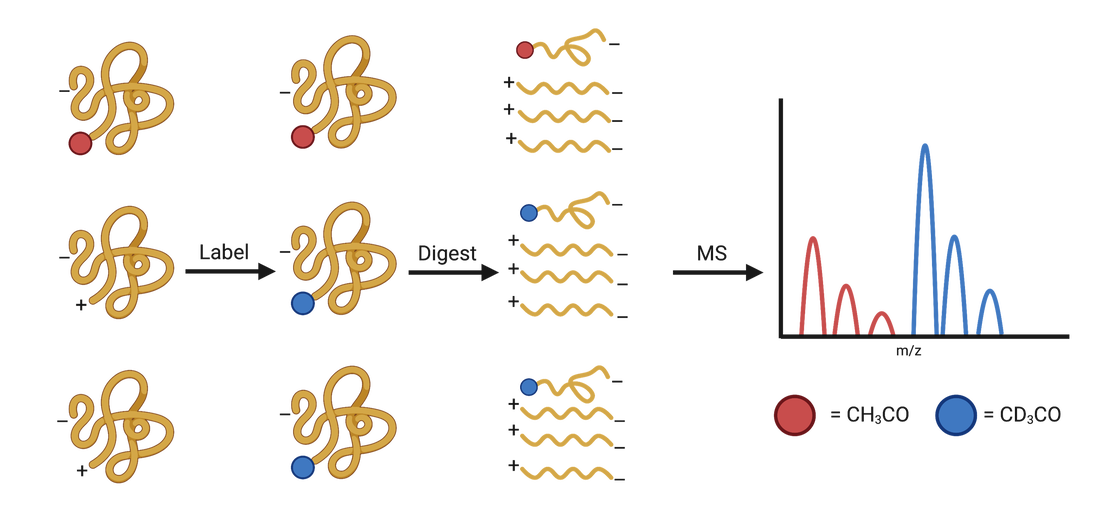
N-terminal acetylation (NTA) is a ubiquitous post-translational modification found in higher organisms. In myco-bacteria, NTA correlates with patho-genesis. We develop novel chemical labeling methods combined with unique instrument implementations to better quantify NTA across mycobacterial samples with varying growth conditions and genetic mutations, to better understand these biological pathways.
|
Data Analysis and Informatics
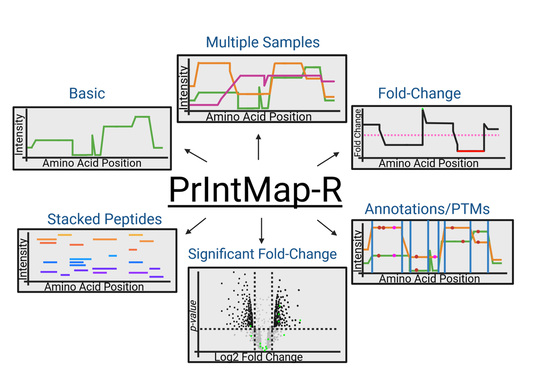
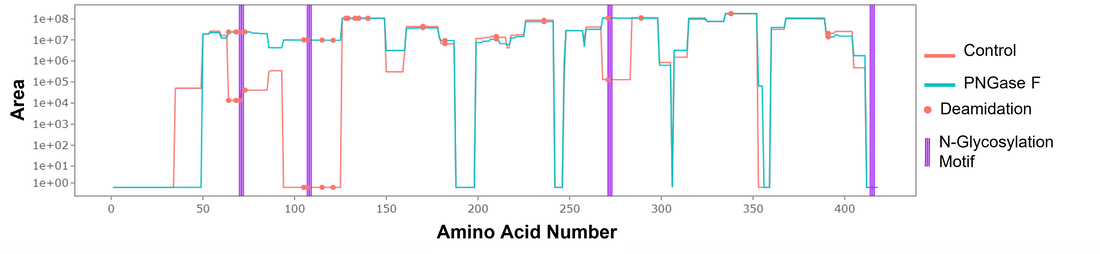
Protein Intensity Mapper (PrIntMap-R)BUP produces rich data that requires programming to analyze. We have developed Protein Intensity Mapper (PrIntMap-R) to produce intensity driven visualizations of specific proteins. This application is used for experimental optimization, modification annotation, and sample comparisons. An example output from PrIntMap-R is shown below, demonstrating one potential use as a tool to help identify N-glycosylation sites. PrIntMap-R can be accessed here.
|
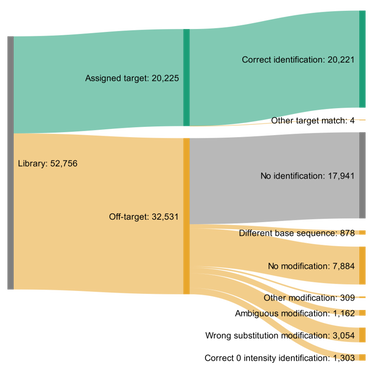
Substitution DetectionWe also push the boundaries of peptide identification past genome-defined sequences. This includes identifying post-translational modifications to amino acids, using transcriptome-based peptide identification, and identifying substitution errors. To the right is a flowchart showing success and failure in identifying peptides that have a substitution error.
|
Capillary Separations
cIEF-MS Interface
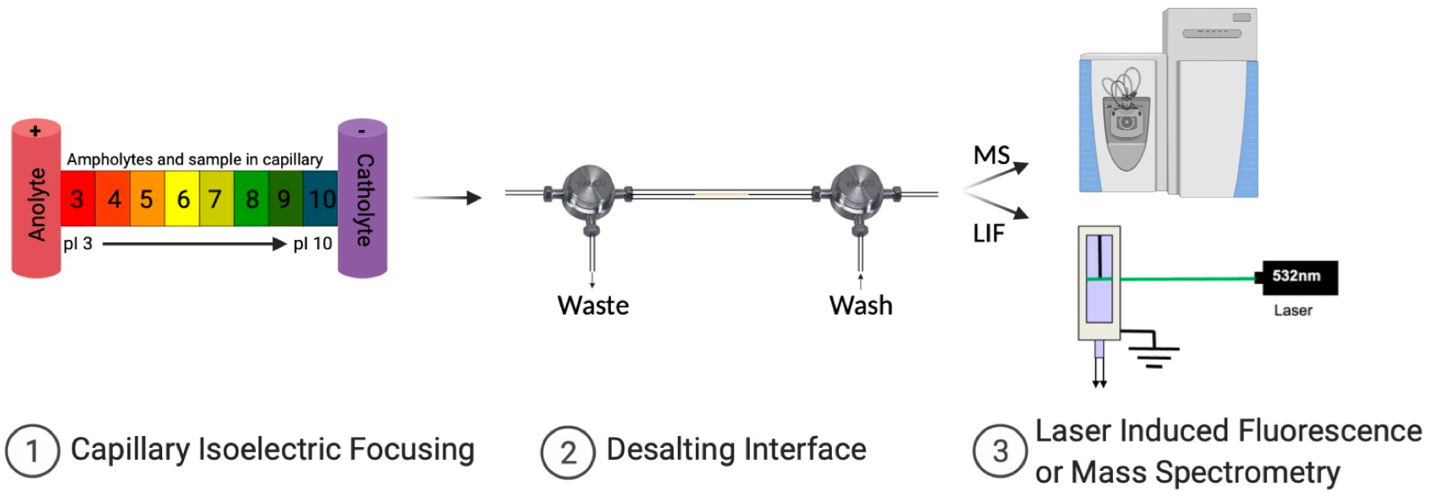
Monoclonal antibodies are an important class of proteins and have been one of the fastest growing therapeutic drug classes. However, the heterogeneity of these therapeutic proteins complicates their development and quality control. Isoelectric focusing (IEF), which separates molecules based on their isoelectric points (pI), has been used to separate therapeutic proteins. Capillary isoelectric focusing (cIEF) is analogous to IEF but is conducted within a liquid capillary, which reduces the analysis time considerably from gel-based focusing. Integrating cIEF with Mass Spectrometry (MS) detection is ideal for protein mass characterization. However, cIEF requires ampholytes, which are low molecular weight species that span a wide range of pI, for the separation to occur. The concentration of ampholytes for high-resolution cIEF is mutually exclusive with robust electrospray ionization (ESI) MS. To address this challenge, we have developed an in-line desalting interface (2) to reduce the amino acid ampholyte concentration after cIEF (1) and prior to ESI (3). The interface consists of a highly selective, semi-permeable Nafion tubing. After focusing, protein and ampholytes are mobilized by electrophoresis and/or pneumatic pressure through the Nafion block; desalted by membrane-exchange and introduced into the source of an ESI-mass spectrometer or detected using laser induced fluorescence (LIF).
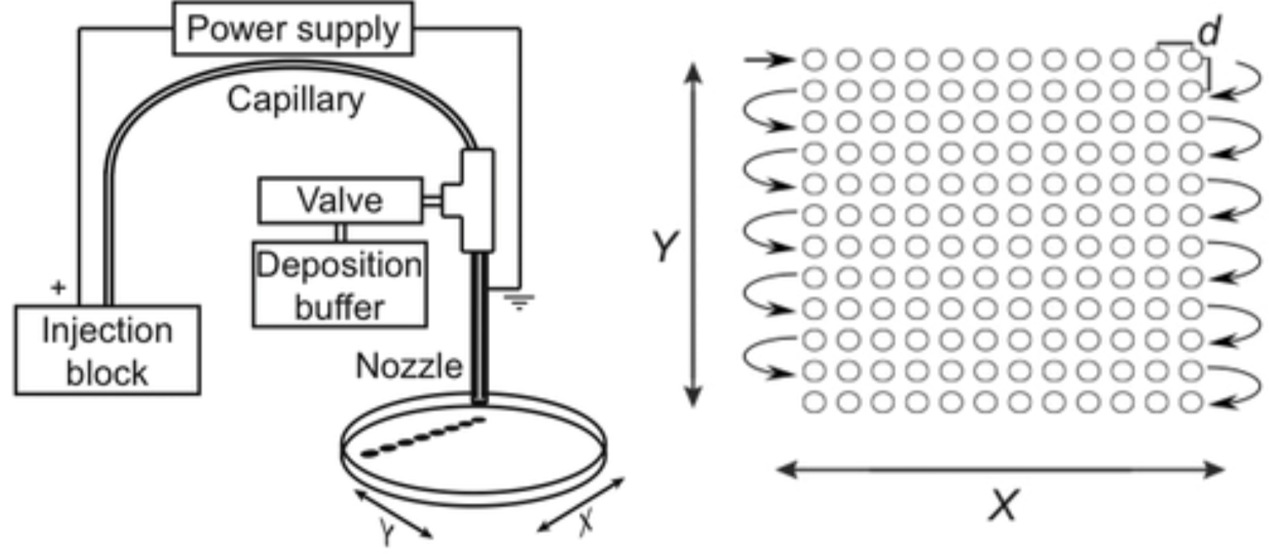
CZE Fraction CollectorWe have developed novel electrophoretic methods to separate and collect complex microbes prior to traditional metagenomic analysis with goals of logarithmic expansion in density and speciation of rare bacteria and viruses.
Components in microorganisms that contribute to their electrophoretic mobility are not fully understood. To address this, we are developing screens to define the genetic requirements that determine electrophoretic mobility in bacteria. |
Some figures created with BioRender